Brandon Sinn Research Page
Select Current Projects
Comparative Genomics
|
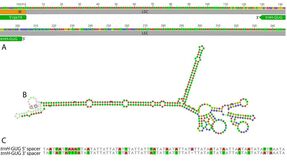
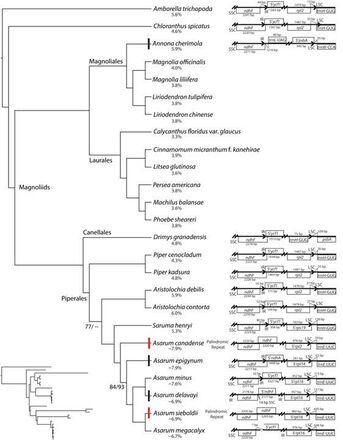
We use genome-scale sequencing and comparative genomic analyses to improve our understanding of the breadth and generation of genomic diversity, and the genetic underpinnings of life history shifts. We recently published the first genomic-scale horizontal gene transfer event between fungal and orchid mitochondrial genomes, which can be found here. Our research of plastid genomes has revealed that the plastomes of the magnoliid genus Asarum contain myriad manifestations of genomic destabilization, such as the only known incorporation of the Small Single Copy region into the Inverted Repeat. This ongoing work will continue to provide more robust insights into plastid genome evolution, stability, and mechanisms underlying genome restructuring.
We are also using RNAseq and Oxford Nanopore techniques to conduct investigations of gene family evolution associated with trophic shifts from autotrophy to mycoheterotropic parasitism. Our model group for this work is the genus Corallorhiza, an early transitional group of mycoheterotrophic orchids. Ongoing analyses of differential gene expression and splicing between partial and fully-mycoheterotrophic Corallorhiza species are revealing pathway and splicing alterations that are associated with this trophic shift, and correlate well with the observed gene losses and structural rearrangements manifest in the plastid genomes of mycoheterotrophic plants. Rachel Muti recently presented our findings on the evolution of WHY1 at Botany 2021. |
Conservation and Population Genetics
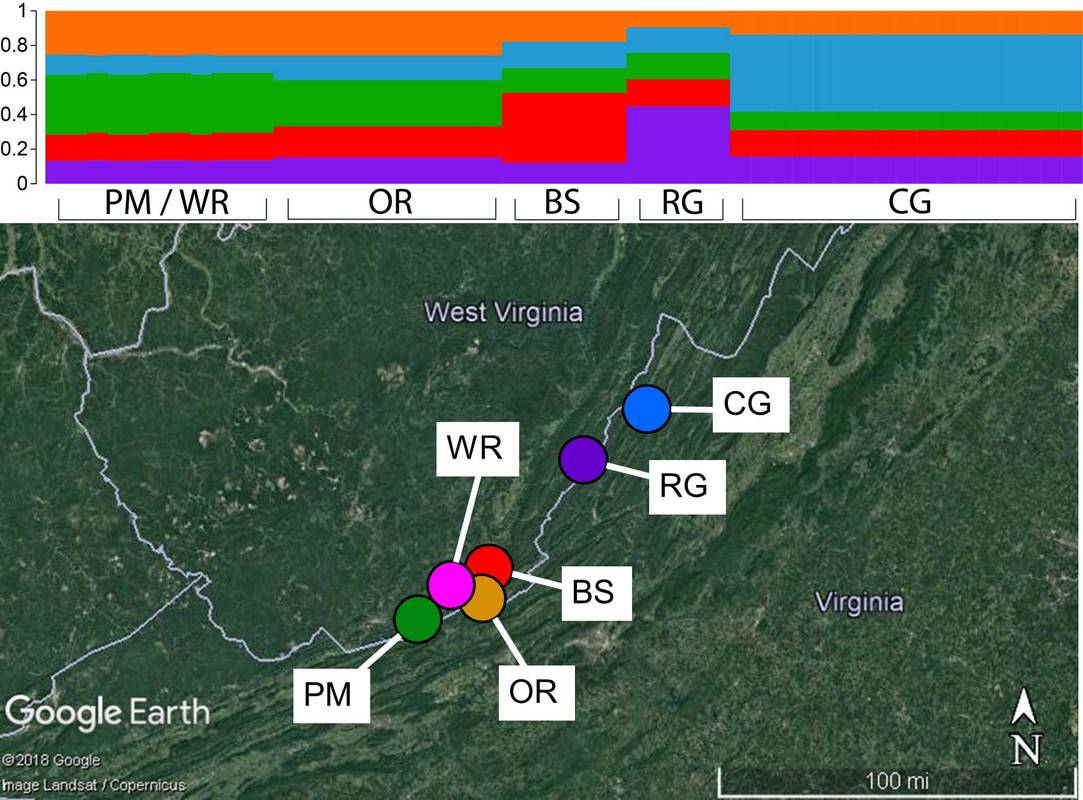
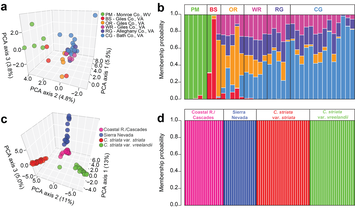
We are currently working to develop novel methods for reduced-representation sequencing of genomes for comparative genomic and population genetic studies involved species of conservation concern. To this end, we are currently collaborating with the Barrett and DiFazio labs at West Virginia University to merge ISSR and genome-scale sequencing techniques to develop a simple, time- and expense- efficient reduced representation protocol that we call ISSRseq. Our publication describing ISSRseq can be found here , and a Git repository, detailed protocol, and wiki for the bioinformatic pipeline are also available. The paper describing this work was recently highlighted as one of eight shortlisted for the Robert May Prize from the British Ecological Society -- you can check out a nice blog post by Sandy Simon (congrats, Sandy!) here. We recently used our method to generate a panel of >47,000 SNPs with which to investigate the population genetics of Corallorhiza bentleyi, a fully-mycoheterotrophic orchid endemic to a handful of counties along the Virginia/West Virginia border. This species is currently IUCN listed, but is not federally protected. We additionally tested ISSRseq using the Corallorhiza striata complex, where ISSRseq resulted in the discovery of >25,000 SNPs spanning potential species boundaries. We next plan to use ISSRseq to characterize the standing genetic diversity in Asarum rosei, a species named by Dr. Sinn in 2017 which is known from only two populations in North Carolina.
Phylogenetic Systematics
|
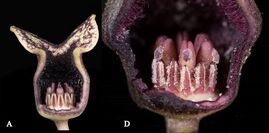
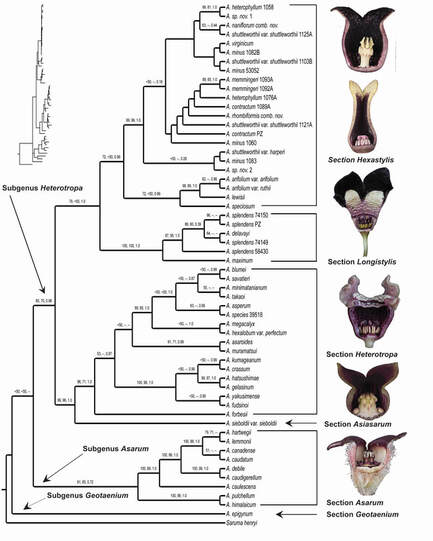
Much of our research involves the generation of phylogenetic hypotheses upon which to found investigations of the dynamics of evolutionary processes. Systematic revisions and species descriptions have resulted from our work. To date, our research has uncovered evidence of character-state associated diversification in the genus Asarum. This work has also resulted in a revised infrageneric classification of the genus, and the description of two species endemic to the southeastern United States. Web-friendly versions of taxonomic keys for Asarum are available here. An article in Youngstown State Magazine featured the discovery of Asarum chueyi, a species named in recognition of an undergraduate mentor.
|
Copyright © Brandon T. Sinn. 2018-2023. All rights reserved.